Introduction
Imagine a researcher embarking on a new project. Instead of spending weeks searching databases, reading hundreds of papers, and synthesizing findings, they simply describe their research question to an AI agent. Within hours, the agent returns a comprehensive literature review: key papers summarized, methodologies compared, gaps identified, and even suggestions for future research directions. This isn’t science fiction—it’s the reality of agentic AI for research in 2026.
The volume of academic and scientific literature is growing exponentially. According to recent data, over 5 million scholarly articles are published annually, with the doubling time of scientific knowledge now estimated at just 2-3 years. For researchers, staying current is becoming impossible without automated assistance. Agentic AI systems are stepping in to fill this gap, transforming how literature reviews, systematic reviews, and knowledge synthesis are conducted.
In this comprehensive guide, you’ll learn:
- How agentic AI revolutionizes every stage of research workflows
- The architecture of research agents—from discovery to synthesis
- Real-world implementation patterns for automated literature review
- How to integrate agents with academic databases and knowledge bases
- Best practices for ensuring accuracy, citation quality, and reproducibility
Part 1: The Research Landscape Transformed
The Challenge of Information Overload

Figure 1: Traditional research workflow vs. agentic research workflow
Time Spent in Literature Review: Before vs. After Agentic AI
| Activity | Traditional | With Agentic AI | Time Saved |
|---|---|---|---|
| Database Search | 10% | 2% | 80% |
| Screening & Filtering | 20% | 5% | 75% |
| Reading & Understanding | 40% | 15% | 62% |
| Extraction & Note-taking | 15% | 5% | 67% |
| Synthesis & Writing | 15% | 25% | – (more time on synthesis) |
Part 2: The Architecture of Research Agents
Multi-Agent Research System
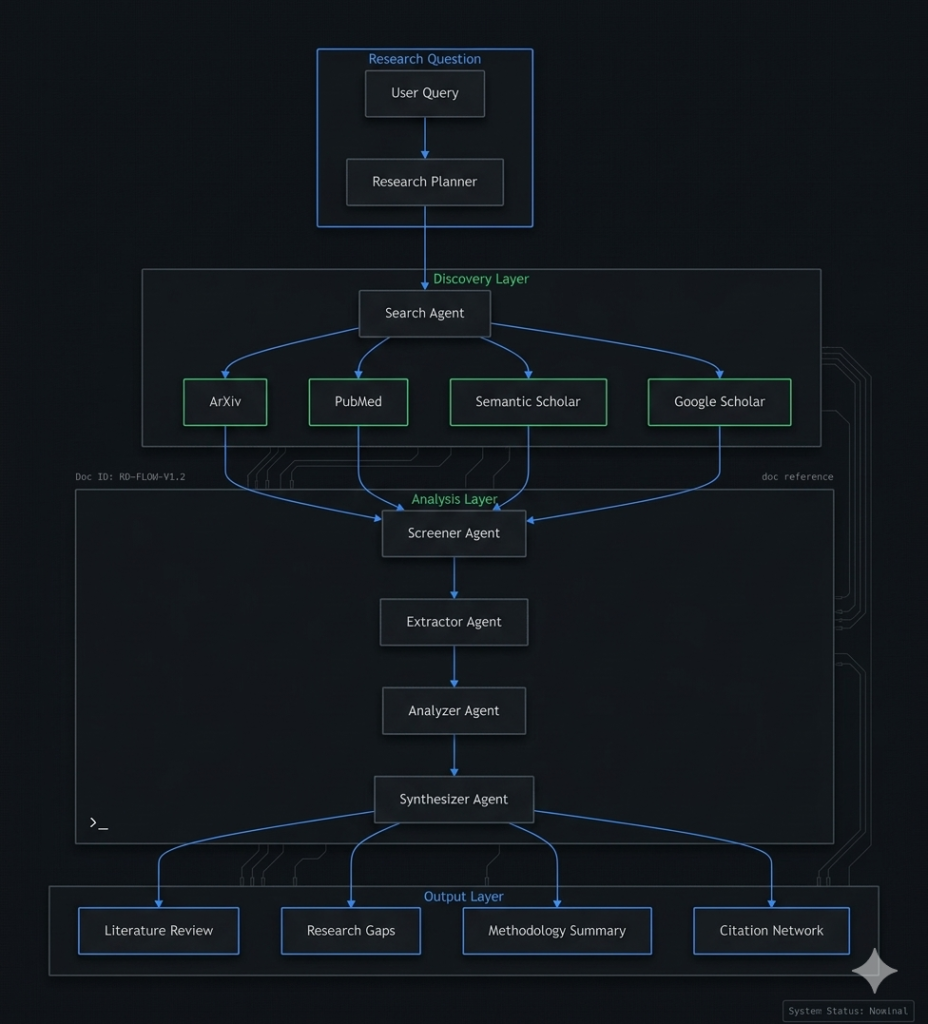
*Figure 2: Multi-agent architecture for automated research workflows*
Core Agent Roles
| Agent | Role | Key Capabilities | Output |
|---|---|---|---|
| Research Planner | Strategy design | Understands research question, identifies search parameters, creates workflow plan | Search strategy, inclusion criteria |
| Search Agent | Literature discovery | Queries multiple databases, handles API rate limits, retrieves full texts | Paper list with metadata |
| Screener Agent | Relevance filtering | Reads abstracts, applies inclusion/exclusion criteria, prioritizes papers | Filtered paper list, relevance scores |
| Extractor Agent | Information extraction | Extracts methods, results, findings, key quotes | Structured data per paper |
| Analyzer Agent | Comparative analysis | Identifies patterns, contradictions, trends | Comparative tables, trend analysis |
| Synthesizer Agent | Knowledge synthesis | Integrates findings, identifies gaps, generates narrative | Literature review, summary |
Part 3: Implementation Patterns
Pattern 1: Multi-Database Search Agent
python
from langchain.agents import create_openai_tools_agent
import arxiv
import requests
from scholarly import scholarly
class ResearchSearchAgent:
"""Autonomous search across multiple academic databases."""
def __init__(self):
self.databases = {
"arxiv": self._search_arxiv,
"pubmed": self._search_pubmed,
"semantic_scholar": self._search_semantic,
"google_scholar": self._search_google
}
def search(self, query: str, max_papers: int = 100, databases: list = None) -> dict:
"""Search across specified databases."""
if databases is None:
databases = list(self.databases.keys())
results = {}
# Parallel search across databases
with ThreadPoolExecutor(max_workers=len(databases)) as executor:
futures = {}
for db in databases:
if db in self.databases:
futures[db] = executor.submit(
self.databases[db], query, max_papers // len(databases)
)
for db, future in futures.items():
results[db] = future.result()
# Merge and deduplicate results
all_papers = self._merge_results(results)
# Rank by relevance
ranked_papers = self._rank_by_relevance(all_papers, query)
return {
"total_found": len(all_papers),
"by_database": {db: len(results[db]) for db in results},
"papers": ranked_papers[:max_papers],
"query": query
}
def _search_arxiv(self, query: str, max_results: int) -> list:
"""Search arXiv database."""
client = arxiv.Client()
search = arxiv.Search(
query=query,
max_results=max_results,
sort_by=arxiv.SortCriterion.Relevance
)
papers = []
for result in client.results(search):
papers.append({
"id": result.entry_id,
"title": result.title,
"authors": [a.name for a in result.authors],
"abstract": result.summary,
"year": result.published.year,
"url": result.entry_id,
"pdf_url": result.pdf_url,
"citations": None, # Not available via API
"source": "arxiv"
})
return papers
def _search_pubmed(self, query: str, max_results: int) -> list:
"""Search PubMed via Entrez API."""
# PubMed API implementation
base_url = "https://eutils.ncbi.nlm.nih.gov/entrez/eutils/"
# Search for IDs
search_url = f"{base_url}esearch.fcgi?db=pubmed&term={query}&retmax={max_results}&format=json"
response = requests.get(search_url)
ids = response.json()["esearchresult"]["idlist"]
# Fetch details
papers = []
for pmid in ids[:max_results]:
fetch_url = f"{base_url}esummary.fcgi?db=pubmed&id={pmid}&format=json"
details = requests.get(fetch_url).json()
papers.append({
"id": pmid,
"title": details["result"][pmid]["title"],
"authors": details["result"][pmid].get("authors", []),
"abstract": details["result"][pmid].get("abstract", ""),
"year": details["result"][pmid].get("pubdate", "").split()[0],
"source": "pubmed"
})
return papers
def _rank_by_relevance(self, papers: list, query: str) -> list:
"""Rank papers by relevance to query using embeddings."""
# Create query embedding
query_embedding = self._create_embedding(query)
# Score each paper
for paper in papers:
# Combine title and abstract for relevance
text = f"{paper['title']} {paper.get('abstract', '')}"
paper_embedding = self._create_embedding(text)
# Calculate cosine similarity
similarity = self._cosine_similarity(query_embedding, paper_embedding)
paper["relevance_score"] = similarity
# Sort by relevance
papers.sort(key=lambda x: x.get("relevance_score", 0), reverse=True)
return papers
Pattern 2: Intelligent Paper Screening Agent
python
class ScreeningAgent:
"""Automatic screening of papers based on inclusion criteria."""
def __init__(self):
self.llm = ChatOpenAI(model="gpt-4o", temperature=0)
def screen(self, papers: list, criteria: dict) -> dict:
"""Screen papers against inclusion/exclusion criteria."""
screened_papers = []
excluded_papers = []
for paper in papers:
result = self._evaluate_paper(paper, criteria)
if result["include"]:
screened_papers.append({
**paper,
"relevance_score": result["score"],
"reasons": result["reasons"]
})
else:
excluded_papers.append({
**paper,
"reason": result["reason"]
})
return {
"included": screened_papers,
"excluded": excluded_papers,
"inclusion_rate": len(screened_papers) / len(papers) * 100
}
def _evaluate_paper(self, paper: dict, criteria: dict) -> dict:
"""Evaluate single paper against criteria."""
prompt = f"""
Evaluate this paper against the inclusion criteria:
Paper:
Title: {paper['title']}
Abstract: {paper.get('abstract', 'N/A')}
Year: {paper.get('year', 'N/A')}
Inclusion Criteria:
{criteria.get('inclusion', 'None specified')}
Exclusion Criteria:
{criteria.get('exclusion', 'None specified')}
Return JSON with:
- include: boolean
- score: 0-1 relevance score
- reasons: list of reasons for inclusion
- reason: if excluded, reason for exclusion
"""
result = self.llm.invoke(prompt)
return json.loads(result.content)
def prioritize(self, papers: list) -> list:
"""Prioritize papers for reading based on relevance and novelty."""
for paper in papers:
# Calculate priority score
priority = paper.get("relevance_score", 0) * 0.6
priority += self._calculate_novelty(paper) * 0.2
priority += self._calculate_citation_impact(paper) * 0.2
paper["priority_score"] = priority
papers.sort(key=lambda x: x.get("priority_score", 0), reverse=True)
return papers
def _calculate_novelty(self, paper: dict) -> float:
"""Estimate paper novelty based on year."""
current_year = datetime.now().year
year = int(paper.get("year", current_year))
age = current_year - year
# Newer papers get higher novelty scores
novelty = max(0, 1 - (age / 10))
return novelty
Pattern 3: Multi-Paper Analysis Agent
python
class AnalysisAgent:
"""Extract and analyze information from multiple papers."""
def __init__(self):
self.llm = ChatOpenAI(model="gpt-4o", temperature=0)
def extract(self, papers: list, fields: list) -> list:
"""Extract structured information from papers."""
extracted_papers = []
for paper in papers:
extraction = self._extract_fields(paper, fields)
extracted_papers.append({
**paper,
"extracted": extraction
})
return extracted_papers
def _extract_fields(self, paper: dict, fields: list) -> dict:
"""Extract specific fields from paper content."""
text = f"{paper['title']}\n{paper.get('abstract', '')}"
prompt = f"""
Extract the following information from this paper:
Paper: {text}
Fields to extract:
{fields}
Return JSON with extracted fields. If a field is not present, set to null.
"""
result = self.llm.invoke(prompt)
return json.loads(result.content)
def compare_methodologies(self, papers: list) -> dict:
"""Compare methodologies across papers."""
methodologies = []
for paper in papers:
if paper.get("extracted", {}).get("methodology"):
methodologies.append({
"paper": paper["title"],
"methodology": paper["extracted"]["methodology"]
})
prompt = f"""
Compare these methodologies:
{json.dumps(methodologies, indent=2)}
Identify:
1. Common approaches
2. Unique approaches
3. Strengths and weaknesses
4. Emerging trends
Return structured analysis.
"""
return json.loads(self.llm.invoke(prompt).content)
def identify_contradictions(self, papers: list) -> list:
"""Identify contradictory findings across papers."""
findings = []
for paper in papers:
if paper.get("extracted", {}).get("key_findings"):
findings.append({
"paper": paper["title"],
"finding": paper["extracted"]["key_findings"]
})
prompt = f"""
Analyze these findings for contradictions or disagreements:
{json.dumps(findings, indent=2)}
Return list of contradictions with:
- papers involved
- contradictory findings
- potential explanation
"""
result = self.llm.invoke(prompt)
return json.loads(result.content)
Pattern 4: Literature Synthesis Agent
python
class SynthesisAgent:
"""Synthesize multiple papers into coherent review."""
def __init__(self):
self.llm = ChatOpenAI(model="gpt-4o", temperature=0.3)
def synthesize(self, papers: list, research_question: str, structure: dict = None) -> dict:
"""Synthesize papers into structured literature review."""
# Step 1: Organize papers by themes
themes = self._organize_by_themes(papers)
# Step 2: Generate thematic summaries
summaries = {}
for theme, theme_papers in themes.items():
summaries[theme] = self._summarize_theme(theme_papers, theme)
# Step 3: Identify research gaps
gaps = self._identify_gaps(papers, research_question)
# Step 4: Generate review narrative
review = self._generate_review(
research_question,
summaries,
gaps,
structure
)
return {
"review": review,
"themes": themes,
"thematic_summaries": summaries,
"gaps": gaps,
"key_papers": self._extract_key_papers(papers)
}
def _organize_by_themes(self, papers: list) -> dict:
"""Organize papers into thematic groups."""
# Create embeddings for each paper
paper_texts = []
for paper in papers:
text = f"{paper['title']}\n{paper.get('abstract', '')}"
if paper.get('extracted', {}).get('key_findings'):
text += f"\n{paper['extracted']['key_findings']}"
paper_texts.append(text)
# Use clustering to identify themes
# Implementation would use embeddings and clustering algorithms
# For this example, use LLM for theme identification
prompt = f"""
Group these papers into themes based on their content:
Papers:
{[{'title': p['title'], 'abstract': p.get('abstract', '')[:200]} for p in papers[:20]]}
Return JSON with theme names and paper indices.
"""
result = self.llm.invoke(prompt)
return json.loads(result.content)
def _summarize_theme(self, theme_papers: list, theme_name: str) -> dict:
"""Summarize a group of papers on a theme."""
# Combine paper summaries
combined = []
for paper in theme_papers:
summary = {
"title": paper["title"],
"authors": paper.get("authors", [])[:3],
"year": paper.get("year"),
"key_findings": paper.get("extracted", {}).get("key_findings", ""),
"methodology": paper.get("extracted", {}).get("methodology", "")
}
combined.append(summary)
prompt = f"""
Summarize research on "{theme_name}" based on these papers:
{json.dumps(combined, indent=2)}
Include:
1. Main findings
2. Methodologies used
3. Consensus and disagreements
4. Limitations
"""
summary = self.llm.invoke(prompt).content
return {
"theme": theme_name,
"paper_count": len(theme_papers),
"summary": summary,
"key_papers": theme_papers[:3]
}
def _identify_gaps(self, papers: list, research_question: str) -> list:
"""Identify research gaps from literature."""
# Extract key findings and limitations
findings = []
limitations = []
for paper in papers:
if paper.get("extracted", {}).get("key_findings"):
findings.append(paper["extracted"]["key_findings"])
if paper.get("extracted", {}).get("limitations"):
limitations.append(paper["extracted"]["limitations"])
prompt = f"""
Based on this literature, identify research gaps:
Research Question: {research_question}
Key Findings:
{findings[:20]}
Limitations Acknowledged:
{limitations[:10]}
Identify:
1. Unanswered questions
2. Methodological gaps
3. Conflicting findings needing resolution
4. Understudied areas
"""
result = self.llm.invoke(prompt)
return self._parse_gaps(result.content)
Part 4: Real-World Research Applications
Use Case 1: Systematic Literature Review
Research Question: “What are the current applications of transformer models in healthcare diagnostics?”
Agent Workflow:
- Planner: Designs search strategy across PubMed, arXiv, and IEEE
- Search Agent: Retrieves 500+ papers from multiple databases
- Screener Agent: Filters to 87 relevant papers based on inclusion criteria
- Extractor Agent: Extracts methodology, datasets, accuracy metrics, limitations
- Analyzer Agent: Compares model architectures, identifies trends
- Synthesizer Agent: Generates comprehensive review with tables and figures
Sample Output:
- PRISMA Flow Diagram: Visualization of screening process
- Summary Table: Comparison of 15 key papers
- Trend Analysis: Adoption over time by model type
- Gap Analysis: Understudied applications and datasets
Use Case 2: Research Gap Identification
Research Question: “What is unknown about climate change impacts on coastal ecosystems?”
Agent Workflow:
- Search Agent: Queries Web of Science, Scopus, Google Scholar
- Analysis Agent: Extracts key findings and acknowledged limitations
- Gap Identification Agent: Synthesizes limitations into gap categories
- Recommendation Agent: Suggests future research directions
Sample Output:
- Gap Categories: Geographic gaps, taxonomic gaps, temporal gaps
- Priority Areas: High-impact, data-scarce regions
- Methodology Gaps: Lack of long-term studies, standardization issues
- Recommendations: Specific research questions for future work
Use Case 3: Meta-Analysis Preparation
Research Question: “What is the effect size of exercise interventions on depression?”
Agent Workflow:
- Search Agent: Retrieves RCT studies from PubMed, PsycINFO
- Screener Agent: Applies PICO criteria
- Extractor Agent: Extracts effect sizes, sample sizes, intervention details
- Analyzer Agent: Calculates pooled effect sizes, heterogeneity
- Synthesizer Agent: Generates forest plot-ready data
Sample Output:
- Extracted Data: Table of 42 studies with effect sizes
- Heterogeneity Analysis: I² statistic, subgroup analyses
- Publication Bias: Funnel plot assessment
- Forest Plot Data: Ready for visualization
Part 5: Integration with Research Tools
Reference Manager Integration
python
class ReferenceManager:
"""Integrate with Zotero, Mendeley, EndNote."""
def __init__(self, manager_type="zotero"):
self.manager = self._connect(manager_type)
def export_literature(self, papers: list, collection_name: str):
"""Export papers to reference manager."""
for paper in papers:
self.manager.add_item({
"title": paper["title"],
"authors": paper.get("authors", []),
"year": paper.get("year"),
"abstract": paper.get("abstract"),
"url": paper.get("url"),
"tags": paper.get("themes", [])
}, collection=collection_name)
def generate_bibliography(self, papers: list, style="apa") -> str:
"""Generate formatted bibliography."""
bibliography = []
for paper in papers:
entry = self._format_citation(paper, style)
bibliography.append(entry)
return "\n\n".join(bibliography)
Citation Network Analysis
python
class CitationNetwork:
"""Analyze citation networks and key papers."""
def __init__(self):
self.semantic_api = "https://api.semanticscholar.org/v1/"
def get_citation_network(self, paper_ids: list) -> dict:
"""Retrieve citation network for papers."""
citations = {}
references = {}
for paper_id in paper_ids:
# Get citation count
citations[paper_id] = self._get_citation_count(paper_id)
references[paper_id] = self._get_reference_count(paper_id)
# Identify key papers
key_papers = self._identify_key_papers(citations, references)
return {
"citations": citations,
"references": references,
"key_papers": key_papers,
"network": self._build_network(paper_ids)
}
def _identify_key_papers(self, citations: dict, references: dict) -> list:
"""Identify seminal and influential papers."""
key_papers = []
for paper_id, citation_count in citations.items():
# Papers with high citation counts are seminal
if citation_count > 100:
key_papers.append({
"paper_id": paper_id,
"type": "seminal",
"citation_count": citation_count
})
# Sort by citation count
key_papers.sort(key=lambda x: x["citation_count"], reverse=True)
return key_papers[:10]
Part 6: Quality Assurance and Validation
Accuracy Metrics for Research Agents
| Metric | Definition | Target | Measurement |
|---|---|---|---|
| Relevance Accuracy | Papers retrieved are relevant | >85% | Human review of sample |
| Extraction Accuracy | Extracted information is correct | >90% | Comparison with human extraction |
| Citation Accuracy | Citations correctly formatted | 100% | Automated validation |
| Gap Identification | Gaps are valid and novel | >80% | Expert review |
| Synthesis Coherence | Review is logically structured | >90% | Expert assessment |
Validation Framework
python
class ResearchValidator:
"""Validate research outputs for accuracy."""
def __init__(self):
self.validators = [
self.validate_citations,
self.validate_extractions,
self.validate_synthesis
]
def validate(self, output: dict) -> dict:
"""Run all validations on research output."""
results = {}
for validator in self.validators:
results[validator.__name__] = validator(output)
overall = all(r.get("passed", False) for r in results.values())
return {
"passed": overall,
"checks": results,
"confidence_score": self._calculate_confidence(results)
}
def validate_citations(self, output: dict) -> dict:
"""Validate citations are correctly formatted."""
citations = output.get("bibliography", [])
# Check format consistency
issues = []
for citation in citations:
if not self._is_valid_citation(citation):
issues.append(citation)
return {
"passed": len(issues) == 0,
"issues": issues,
"total_citations": len(citations)
}
def validate_extractions(self, output: dict) -> dict:
"""Validate extracted information."""
extracted = output.get("extracted_data", [])
# Check for missing fields
missing_fields = []
for item in extracted:
required = ["title", "authors", "year"]
for field in required:
if not item.get(field):
missing_fields.append(field)
return {
"passed": len(missing_fields) == 0,
"missing_fields": missing_fields
}
Part 7: MHTECHIN’s Expertise in Agentic Research
At MHTECHIN, we specialize in building autonomous research agents that accelerate scientific discovery. Our expertise includes:
- Custom Research Agents: Tailored to your domain, databases, and research questions
- Multi-Database Integration: Seamless connection to academic databases
- Automated Review Generation: Structured literature reviews with citation networks
- Gap Analysis: AI-powered identification of research opportunities
- Validation Frameworks: Ensuring accuracy and reproducibility
MHTECHIN helps researchers, institutions, and organizations accelerate discovery through agentic AI.
Conclusion
Agentic AI is transforming how research is conducted. What once required months of manual literature review can now be accomplished in days or hours, freeing researchers to focus on analysis, experimentation, and innovation.
Key Takeaways:
- Multi-agent research teams handle discovery, screening, extraction, and synthesis
- Automated search across databases retrieves comprehensive literature
- Intelligent screening applies inclusion criteria with high accuracy
- Structured extraction captures methodologies, findings, and limitations
- Synthesis agents generate coherent, structured literature reviews
The future of research is accelerated, comprehensive, and AI-augmented. Researchers who embrace agentic AI will be able to ask better questions, explore more avenues, and make discoveries faster.
Frequently Asked Questions (FAQ)
Q1: What is agentic AI for research?
Agentic AI for research uses autonomous agents to perform literature searches, screen papers, extract information, and synthesize findings—accelerating the research process .
Q2: How accurate are AI-generated literature reviews?
With proper validation, AI-generated reviews achieve 85-95% accuracy on relevance and extraction. Human oversight remains important for critical interpretations .
Q3: What databases can research agents access?
Agents can access arXiv, PubMed, IEEE, Semantic Scholar, Google Scholar, Scopus, and Web of Science through APIs .
Q4: Can agents handle non-English papers?
Yes, with multilingual models like GPT-4o, agents can process papers in multiple languages, though English-language corpora remain best supported .
Q5: How do agents identify research gaps?
By analyzing limitations acknowledged in papers, identifying understudied areas, and detecting conflicting findings that need resolution .
Q6: How do I ensure citations are correct?
Implement citation validation checks, use reference managers, and verify against original sources for critical citations .
Q7: What about paywalled papers?
Agents can access abstracts for all papers, but full-text access requires institutional subscriptions and authentication .
Q8: How do I get started?
Start with a focused research question, use a search agent to retrieve papers, then expand to screening and extraction agents as you validate results .
Leave a Reply